
go-elite
Software Description
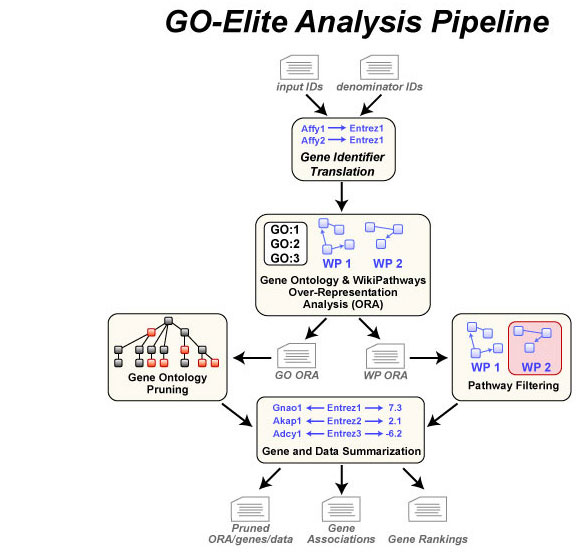
GO-Elite (http://genmapp.org/go_elite) is a software tool designed to identify a minimal non-redundant set of biological Ontology (GO) terms or pathways to describe a particular set of genes or metabolites. Default resources include multiple ontologies (Gene, Disease, Phenotype), pathways (WikiPathways, KEGG, Pathway Commons), putative regulatory targets (transcription, microRNA, domains) and cellular biomarkers.
This tool calculates over-representation statistics similar to the program MAPPFinder and subsequently produces filtered Ontology and pathway results along with corresponding gene annotations and summarized expression data. Multiple options for pathway visualization are also available. This software can be run using an intuitive graphical user interface, through command-line arguments, using a web interface and through the program GenMAPP-CS. Please see our complete documentation for more details.
News
- GO-Elite version 1.2.5 is now available (7-7-12)
- The application note "GO-Elite: A Flexible Solution for Pathway and Ontology Over-Representation" has been published and is now online (6-29-12)
- All news post details can be found here
Project Information
- License: Apache License 2.0
- 6 stars
- svn-based source control
Labels:
Bioinformatics
Biology
Python
Pathway
 Code
Code